How the influenza virus achieves efficient viral RNA replication
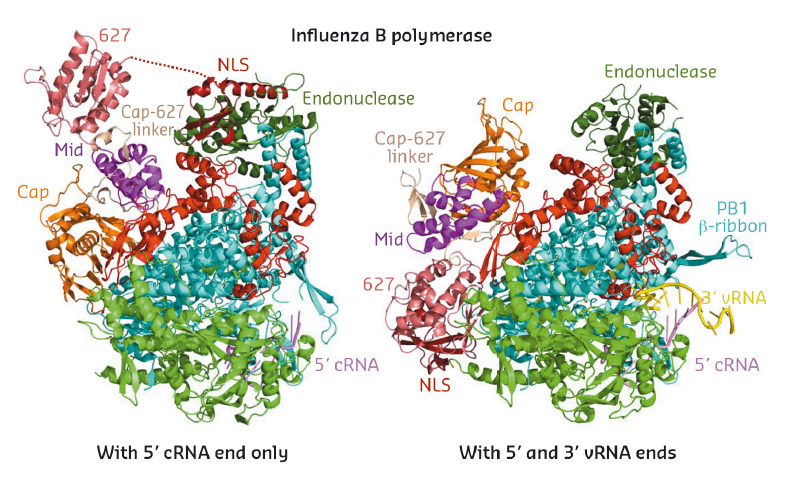
New insights on how subunits of the influenza virus polymerase co-evolve to ensure efficient viral RNA replication are provided by a study published October 3 in the open-access journal PLOS Pathogens by Nadia Naffakh of the Institut Pasteur, and colleagues. As the authors note, the findings could lead to novel strategies for antiviral drug development.
Because of their yearly recurrence and the occasional emergence of pandemics, influenza viruses represent a worldwide major public health threat. Enhancing fundamental knowledge about the influenza RNA-polymerase, which is an enzyme that consists of three subunits (i.e., a heterotrimer) and ensures transcription and replication of the viral genome, is essential to reach the goal of better prevention and treatment of disease.
In the new study, Naffakh and colleagues gained new insights into viral polymerase function. They showed that the polymerase subunits co-evolve to ensure not only optimal inter-subunit cooperation within the heterotrimer, but also proper levels dimerization — the process by which pairs of heterotrimers attach together — which appears to be essential for efficient viral RNA replication. The findings point to influenza polymerase dimerization as a feature that can restrict genetic reassortment, a major evolutionary mechanism in which influenza viruses swap gene segments, and could become an attractive target for antiviral drug development.
Original published by Sciencedaily

(Illustration photo: Influenza B polymerase)
(Source: http://www.esrf.eu/home/UsersAndScience/Publications/Highlights/highlights-2016/SB/SB04.html)