New antibiotics target the outer membrane of bacteria
Antibiotic resistance is a growing global public-health problem. One group of bacteria, called Gram-negative bacteria, is particularly difficult to treat, because the cells are shielded by a double-membrane envelope, which constitutes a formidable barrier to antibiotics. When antibiotics do breach the membranes, these bacteria often use efflux pumps to remove the drugs. Three papers (two in Nature and one in the Proceedings of the National Academy of Sciences) now describe antibiotics that overcome these obstacles by targeting, directly or indirectly, a protein integral to the outer membrane.
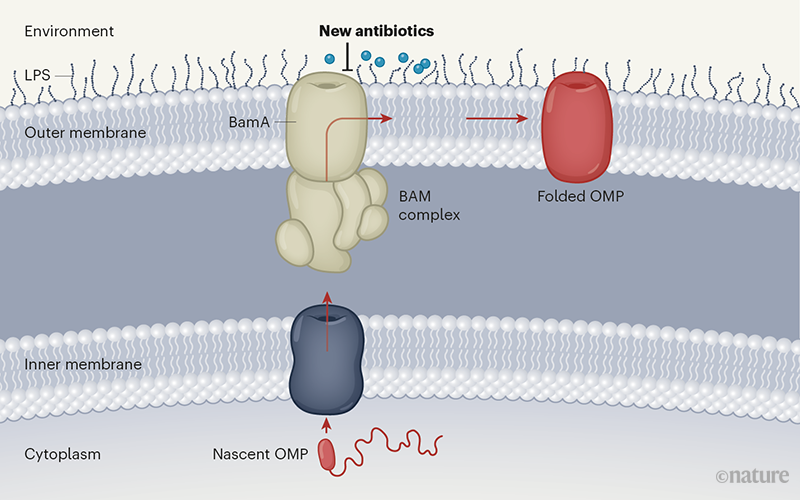
The outer membrane of Gram-negative bacteria contains lipopolysaccharide (LPS) molecules in its outer leaflet, with outer-membrane proteins (OMPs) spanning the entire outer membrane. OMPs are folded into the membrane by a protein complex called the β-barrel assembly machine (BAM), the central component of which, BamA, is an OMP itself. Because BamA is exposed to the extracellular space, it could be an Achilles heel in the bacterial shield — inhibitors that access BamA would not need to penetrate the cell. Indeed, a proof-of-concept study has shown that this approach inhibits OMP folding and compromises membrane integrity, albeit by an unknown mechanism.
Imai et al. turned to Gram-negative bacteria that live symbiotically in the gut of nematode worms and can secrete antibiotics to fend off competing bacteria — including other Gram-negative species. A screen of the secretions from 22 of these symbionts revealed a Gram-negative-targeting antibiotic, which the authors named darobactin.
Darobactin displayed antibiotic activity against multiple Gram-negative bacteria, both in vitro and in infected mice, including against several drug-resistant human pathogens such as polymyxin-resistant Pseudomonas aeruginosa and β-lactam-resistant Klebsiella pneumoniae and Escherichia coli. Darobactin was not toxic to human cells at the concentrations at which it was an effective antibiotic.
The authors provided evidence that darobactin and BamA bind to each other directly, using a technique called isothermal titration calorimetry, which measures the heat changes associated with physical interactions between molecules. The results of nuclear magnetic resonance (NMR) spectroscopy experiments were also consistent with direct binding, and suggested that the antibiotic stabilizes the protein in a potentially inactive conformation.

Overcoming a double-membrane barrier
(Gram-negative bacteria are protected by inner and outer membranes. The outer membrane contains lipopolysaccharide (LPS) molecules in the outer layer and integral outer-membrane proteins (OMPs). These proteins are synthesized in the cell’s cytoplasm and transported to the space between the membranes by the translocation machinery (dark blue). From here, they are captured, inserted and folded into the outer membrane by the BAM protein complex (red arrows). BamA is the central component of BAM and is accessible from the bacterial surface. Three studies describe new antibiotics that seem to target BamA, preventing the normal OMP folding that is required for bacterial survival)
Original published by Nature with slightly modification